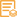主要信息
Target
Histone H3 Di Methyl Lys9
Host Species
Rabbit
Reactivity
Human, Mouse, Rat
Applications
WB
MW
15-17kD (Observed)
Conjugate/Modification
Di Methyl

货号: YH0034
规格
价格
货期
数量
200μL
¥4,680.00
一个月
0
100μL
¥2,800.00
一个月
0
50μL
¥1,500.00
一个月
0
加入购物车


已收藏


收藏
详细信息
推荐稀释比
WB 1:500-1000
组成
PBS, pH 7.4, containing 0.5%BSA, 0.02% sodium azide as Preservative and 50% Glycerol.
特异性
The antibody detects endogenous Histone H3 (Di Methyl Lys9) protein.
纯化工艺
The antibody was affinity-purified from rabbit antiserum by affinity-chromatography using epitope-specific immunogen.
储存
-15°C to -25°C/1 year (Do not lower than -25°C)
实测条带
15-17kD
修饰
Di Methyl
克隆性
Polyclonal
同种型
IgG
相关产品
抗原&靶点信息
免疫原:
Synthetic Peptide of Histone H3 (Di Methyl Lys9)
展开内容
特异性:
The antibody detects endogenous Histone H3 (Di Methyl Lys9) protein.
展开内容
基因名称:
HIST1H3A/HIST1H3B/HIST1H3C/HIST1H3D/HIST1H3E/HIST1H3F/HIST1H3G/HIST1H3H/HIST1H3I/HIST1H3J/HIST2H3A/HIST2H3C/HIST2H3D/H3F3A/H3F3B
展开内容
蛋白名称:
Histone H3.1/Histone H3.2/Histone H3.3
展开内容
别名:
H3K9ME2 ;
HIST1H3A ;
H3FA ;
HIST1H3B ;
H3FL ;
HIST1H3C ;
H3FC ;
HIST1H3D ;
H3FB ;
HIST1H3E ;
H3FD ;
HIST1H3F ;
H3FI ;
HIST1H3G ;
H3FH ;
HIST1H3H ;
H3FK ;
HIST1H3I ;
H3FF ;
HIST1H3J ;
H3FJ ;
Histone H3.1 ;
Histone H3/a ;
Histone H3/b ;
Histone H3/c ;
Histone H3/d ;
Histone H3/f ;
Histone H3/h ;
Histone H3/i ;
Histone H3/j ;
Histone H3/k ;
Histone H3/l ;
HIST2H3A ;
HIST2H3C ;
H3F2 ;
H3FM ;
HIST2H3D ;
Histone H3.2 ;
Histone H3/m ;
Histone H3/o ;
H3F3A ;
H3.3A ;
H3F3 ;
PP781 ;
H3F3B ;
H3.3B ;
Histone H3.3
HIST1H3A ;
H3FA ;
HIST1H3B ;
H3FL ;
HIST1H3C ;
H3FC ;
HIST1H3D ;
H3FB ;
HIST1H3E ;
H3FD ;
HIST1H3F ;
H3FI ;
HIST1H3G ;
H3FH ;
HIST1H3H ;
H3FK ;
HIST1H3I ;
H3FF ;
HIST1H3J ;
H3FJ ;
Histone H3.1 ;
Histone H3/a ;
Histone H3/b ;
Histone H3/c ;
Histone H3/d ;
Histone H3/f ;
Histone H3/h ;
Histone H3/i ;
Histone H3/j ;
Histone H3/k ;
Histone H3/l ;
HIST2H3A ;
HIST2H3C ;
H3F2 ;
H3FM ;
HIST2H3D ;
Histone H3.2 ;
Histone H3/m ;
Histone H3/o ;
H3F3A ;
H3.3A ;
H3F3 ;
PP781 ;
H3F3B ;
H3.3B ;
Histone H3.3
展开内容
数据库链接:
背景:
Histones are basic nuclear proteins that are responsible for the nucleosome structure of the chromosomal fiber in eukaryotes. This structure consists of approximately 146 bp of DNA wrapped around a nucleosome , an octamer composed of pairs of each of the four core histones (H2A , H2B , H3 , and H4) . The chromatin fiber is further compacted through the interaction of a linker histone , H1 , with the DNA between the nucleosomes to form higher order chromatin structures. This gene is intronless and encodes a replication-dependent histone that is a member of the histone H3 family. Transcripts from this gene lack polyA tails; instead , they contain a palindromic termination element. This gene is found in the large histone gene cluster on chromosome 6p22-p21.3. [provided by RefSeq , Aug 2015] ,
展开内容
功能:
Caution:Was originally (PubMed:2587222) thought to originate from mouse. ,developmental stage:Expressed during S phase , then expression strongly decreases as cell division slows down during the process of differentiation. ,Function:Core component of nucleosome. Nucleosomes wrap and compact DNA into chromatin , limiting DNA accessibility to the cellular machineries which require DNA as a template. Histones thereby play a central role in transcription regulation , DNA repair , DNA replication and chromosomal stability. DNA accessibility is regulated via a complex set of post-translational modifications of histones , also called histone code , and nucleosome remodeling. ,mass spectrometry:Monoisotopic with N-acetylserine PubMed:16457589 ,miscellaneous:This histone is only present in mammals and is enriched in acetylation of Lys-15 and dimethylation of Lys-10 (H3K9me2) . ,PTM:Acetylation is generally linked to gene activation. Acetylation on Lys-10 (H3K9ac) impairs methylation at Arg-9 (H3R8sme2) . Acetylation on Lys-19 (H3K18ac) and Lys-24 (H3K24ac) favors methylation at Arg-18 (H3R17me) . ,PTM:Asymmetric dimethylation at Arg-18 (H3R17me2a) by CARM1 is linked to gene activation. Symmetric dimethylation at Arg-9 (H3R8sme2) by PRMT5 is linked to gene repression. Asymmetric dimethylation at Arg-3 (H3R2me2a) by PRMT6 is linked to gene repression and is mutually exclusive with H3 Lys-5 methylation (H3K4me2 and H3K4me3) . H3R2me2a is present at the 3' of genes regardless of their transcription state and is enriched on inactive promoters , while it is absent on active promoters. ,PTM:Citrullination at Arg-9 (H3R8ci) and/or Arg-18 (H3R17ci) by PADI4 impairs methylation and represses transcription. ,PTM:Deiminated on Arg-4 in granulocytes upon calcium entry. ,PTM:Methylation at Lys-5 (H3K4me) , Lys-37 (H3K36me) and Lys-80 (H3K79me) are linked to gene activation. Methylation at Lys-5 (H3K4me) facilitates subsequent acetylation of H3 and H4. Methylation at Lys-80 (H3K79me) is associated with DNA double-strand break (DSB) responses and is a specific target for TP53BP1. Methylation at Lys-10 (H3K9me) and Lys-28 (H3K27me) are linked to gene repression. Methylation at Lys-10 (H3K9me) is a specific target for HP1 proteins (CBX1 , CBX3 and CBX5) and prevents subsequent phosphorylation at Ser-11 (H3S10ph) and acetylation of H3 and H4. Methylation at Lys-5 (H3K4me) and Lys-80 (H3K79me) require preliminary monoubiquitination of H2B at 'Lys-120'. Methylation at Lys-10 (H3K9me) and Lys-28 (H3K27me) are enriched in inactive X chromosome chromatin. ,PTM:Monoubiquitination of Lys-120 by RING1 and RNF2/RING2 complex gives a specific tag for epigenetic transcriptional repression and participates in X chromosome inactivation of female mammals. It is involved in the initiation of both imprinted and random X inactivation. Ubiquitinated H2A is enriched in inactive X chromosome chromatin. Ubiquitination of H2A functions downstream of methylation of 'Lys-27' of histone H3. Monoubiquitination of Lys-120 by RNF2/RING2 can also be induced by ultraviolet and may be involved in DNA repair. Following DNA double-strand breaks (DSBs) , it is ubiquitinated through 'Lys-63' linkage of ubiquitin moieties by the E2 ligase UBE2N and the E3 ligases RNF8 and RNF168 , leading to the recruitment of repair proteins to sites of DNA damage. Monoubiquitination and ionizing radiation-induced 'Lys-63'-linked ubiquitination are distinct events. ,PTM:Phosphorylated at Thr-4 (H3T3ph) by GSG2/haspin during prophase and dephosphorylated during anaphase. At centromeres , specifically phosphorylated at Thr-12 (H3T11ph) from prophase to early anaphase , probably by DAPK3 (By similarity) . Phosphorylation at Ser-11 (H3S10ph) by AURKB is crucial for chromosome condensation and cell-cycle progression during mitosis and meiosis. In addition phosphorylation at Ser-11 (H3S10ph) by RPS6KA4 and RPS6KA5 is important during interphase because it enables the transcription of genes following external stimulation , like mitogens , stress , growth factors or UV irradiation and result in the activation of genes , such as c-fos and c-jun. Phosphorylation at Ser-11 , which is linked to gene activation , prevents methylation at Lys-10 (H3K9me) but facilitates acetylation of H3 and H4. Phosphorylation at Ser-11 (H3S10ph) by AURKB mediates the dissociation of HP1 proteins (CBX1 , CBX3 and CBX5) from heterochromatin. Phosphorylation at Ser-11 (H3S10ph) is also an essential regulatory mechanism for neoplastic cell transformation. Phosphorylated at Ser-29 (H3S28ph) by MLTK isoform 1 , RPS6KA5 or AURKB during mitosis or upon ultraviolet B irradiation. ,PTM:Phosphorylated at Thr-4 (H3T3ph) by GSG2/haspin during prophase and dephosphorylated during anaphase. At centromeres , specifically phosphorylated at Thr-12 (H3T11ph) from prophase to early anaphase , probably by DAPK3. Phosphorylation at Ser-11 (H3S10ph) by AURKB is crucial for chromosome condensation and cell-cycle progression during mitosis and meiosis. In addition phosphorylation at Ser-11 (H3S10ph) by RPS6KA4 and RPS6KA5 is important during interphase because it enables the transcription of genes following external stimulation , like mitogens , stress , growth factors or UV irradiation and result in the activation of genes , such as c-fos and c-jun. Phosphorylation at Ser-11 (H3S10ph) , which is linked to gene activation , prevents methylation at Lys-10 (H3K9me) but facilitates acetylation of H3 and H4. Phosphorylation at Ser-11 (H3S10ph) by AURKB mediates the dissociation of HP1 proteins (CBX1 , CBX3 and CBX5) from heterochromatin. Phosphorylation at Ser-11 (H3S10ph) is also an essential regulatory mechanism for neoplastic cell transformation. Phosphorylated at Ser-29 by MLTK isoform 1 , RPS6KA5 or AURKB during mitosis or upon ultraviolet B irradiation. ,PTM:Phosphorylation on Ser-2 is enhanced during mitosis. Phosphorylation on Ser-2 by RPS6KA5/MSK1 directly represses transcription. Acetylation of H3 inhibits Ser-2 phosphorylation by RPS6KA5/MSK1. ,PTM:Symmetric dimethylation on Arg-4 by the PRDM1/PRMT5 complex may play a crucial role in the germ-cell lineage. ,PTM:The chromatin-associated form is phosphorylated on Thr-121 during mitosis. ,PTM:Ubiquitinated by the CUL4-DDB-RBX1 complex in response to ultraviolet irradiation. This may weaken the interaction between histones and DNA and facilitate DNA accessibility to repair proteins. ,similarity:Belongs to the histone H2A family. ,similarity:Belongs to the histone H3 family. ,subunit:The nucleosome is a histone octamer containing two molecules each of H2A , H2B , H3 and H4 assembled in one H3-H4 heterotetramer and two H2A-H2B heterodimers. The octamer wraps approximately 147 bp of DNA. ,subunit:The nucleosome is a histone octamer containing two molecules each of H2A , H2B , H3 and H4 assembled in one H3-H4 heterotetramer and two H2A-H2B heterodimers. The octamer wraps approximately 147 bp of DNA. During nucleosome assembly the chaperone ASF1A interacts with the histone H3-H4 heterodimer. ,
展开内容
细胞定位:
Nucleus. Chromosome.
展开内容
研究领域:
>>Neutrophil extracellular trap formation ;
>>Alcoholism ;
>>Shigellosis ;
>>Transcriptional misregulation in cancer ;
>>Systemic lupus erythematosus
>>Alcoholism ;
>>Shigellosis ;
>>Transcriptional misregulation in cancer ;
>>Systemic lupus erythematosus
展开内容
信号通路
文献引用({{totalcount}})
货号: YH0034
规格
价格
货期
数量
200μL
¥4,680.00
一个月
0
100μL
¥2,800.00
一个月
0
50μL
¥1,500.00
一个月
0
加入购物车


已收藏


收藏
Recently Viewed Products
Clear allToggle night Mode
{{pinfoXq.title || ''}}
Catalog: {{pinfoXq.catalog || ''}}
Filter:
All
{{item.name}}
{{pinfo.title}}
-{{pinfo.catalog}}
主要信息
Target
{{pinfo.target}}
Reactivity
{{pinfo.react}}
Applications
{{pinfo.applicat}}
Conjugate/Modification
{{pinfo.coupling}}/{{pinfo.modific}}
MW (kDa)
{{pinfo.mwcalc}}
Host Species
{{pinfo.hostspec}}
Isotype
{{pinfo.isotype}}
产品 {{index}}/{{pcount}}
上一个产品
下一个产品
{{pvTitle}}
滚轮缩放图片
{{pvDescr}}